Bioinformatics Assignment on Caspase-8
| ✓ Paper Type: Free Assignment | ✓ Study Level: University / Undergraduate |
| ✓ Wordcount: 956 words | ✓ Published: 12 Oct 2021 |
Genomic location, protein function & domains/structural features 281
The gene encoding caspase-8, an apoptosis-related cysteine peptidase, is located on chromosome 2 in the human genome. As a member of the cysteine-aspartic acid protease family it has roles in apoptosis, macrophage differentiation, and heart development. Caspase8 exists as an inactive proenzyme consisting of an N-terminal prodomain (including two death effector domains (DED)) as well as a large protease subunit containing catalytic sites and a small protease subunit.
Casp8 is broadly expressed in bone marrow, lymph node, and 21 other tissues. It is an initiator caspase that acts upon executioner caspases such as caspase- 3 (Wang, et al., 2010). Once activated it uses the SH group of a Cys side chain to catalyze cleavage of peptide bonds at aspartyl residues in the substrate. Casp8 also possesses an allosteric binding site aiding its function and acts at neutral pH. The subsite preferences of Casp8 are Asp in P1, Glu in P3, and small residues in P1′ (Stennicke, et al., 2000).
Ca8 is vital in regulating and initiating death receptor-mediated activation of apoptosis (Tummers & Green, 2017). Its role in programmed cell death is induced by Fas, a ligand that binds to Fas receptors on the cell surface. The caspase is activated by proteolytic processing at internal Asp residues. The FADD-like DED may interact with FADD, a Fas-interacting adaptor protein. The formation of a death-inducing signaling complex, including Casp8 and FADD is essential for extrinsic apoptosis. FADD binding to the receptor induces a conformational change exposing the DED to bind Casp8 resulting in a fully matured, active Casp8 which cleaves the zymogens of effector caspases, like caspase-3 between their large and small subunits thus activating them. Casp8 also cleaves BID, activating BAX and BAK, which mediate MOMP. This ultimately allows executioner caspases to promote apoptosis.
Caspase 8 is composed of the following domains and structural features identified by InterProScan (Blum, et al., 2020): 117
– DED domain, at position 1-179, a homotypic protein interaction module composed of a six-helical bundle involved in the interaction between procaspase-8 and FADD DEDs and mediates homodimerization of the proenzyme. Cleavage of an aspartate connecting the DED to the large subunit results in the release of the fully matured enzyme and initiation of apoptosis.
– Peptidase C14A domain (225-374) consisting of two caspase catalytic subunits derived from the p45 precursor:
– p18 (217 – 374), consisting of:
– a histidine active site (304 – 318)
– a cysteine active site (351 – 362)
– p10 (385-478)
The key catalytic residues located on p18, H317, and C360 comprise the catalytic triad along with the backbone carbonyl oxygen group of Arg358.
 Figure 1(a) and (b): Red – peptidase p18 subunit, blue – peptidase p10 subunit, yellow – histidine active site, pink – cysteine active site. (PDB ID: 3KJQ) (A) A ribbon representation shows that the protein is heterodimeric and composed of a 3-layered alpha/beta/alpha sandwich. (B) A surface view of caspase 8 shows the inhibitor covalently bound via the histidine and cysteine active sites indicating their location in the protein (The PyMOL Molecular Graphics System, 2021; Drozdetskiy, et al., 2015).
Figure 1(a) and (b): Red – peptidase p18 subunit, blue – peptidase p10 subunit, yellow – histidine active site, pink – cysteine active site. (PDB ID: 3KJQ) (A) A ribbon representation shows that the protein is heterodimeric and composed of a 3-layered alpha/beta/alpha sandwich. (B) A surface view of caspase 8 shows the inhibitor covalently bound via the histidine and cysteine active sites indicating their location in the protein (The PyMOL Molecular Graphics System, 2021; Drozdetskiy, et al., 2015).
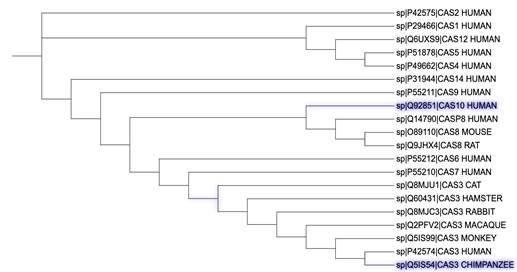
Figure 2: Protein sequence-based phylogenetic tree of cysteine-aspartic acid protease family caspases. This tree was generated using PSI-BLAST, ClustalO, and iTOL based on sequence alignment of the amino acid sequences for caspases by the neighbor-joining distance-based method (Altschul, et al., 1997; Letunic & Bork, 2019).
The tree shows how closely, and distantly related caspases are across different species.
The Evolutionary History of the cysteine-aspartic acid protease gene family: 178
Caspase-8 belongs to the highly conserved gene family of cysteine-aspartic acid caspases which recognize tetrapeptide motifs and cleave peptide bonds after aspartate residues with roles in apoptosis, cell differentiation, and inflammation (Sakamaki, et al., 2014). Despite the distinct number of caspases for each species e.g. 11 caspases in humans, 14 in rodents and cows. Their role in apoptosis may be an evolutionarily conserved function (Sakamaki, et al., 2009). Many of these caspases are orthologues to caspases in other eukaryotic species, as shown in figure 2, suggesting that through evolution the function of caspases has been conserved.
Apoptotic caspases may have evolved from an ancestral immune system into the two subfamilies of initiators and effectors (Grinshpon, et al., 2019). Due to the different number of caspases in each organism, caspases likely diverged over evolutionary time, through gene duplication events, becoming species-specific, and undergoing neo-functionalism.
In figure 2, the initiator caspases with two DEDs – caspase-8 and -10, the executioner caspases-3, -6, and -7 with a short prodomain and inflammatory caspases-4, -5, -1, and -12 with a conserved long prodomain are clustered together. This shows that caspase function is conserved across different species.
How deletions, SNPs and/or copy number variations in the caspase-8 gene are relevant to the associated disorders: 168
Caspase-8 is associated with several genetic disorders such as lung cancer, hepatocellular carcinoma, and breast cancer. An autosomal recessive mutation in the Casp8 gene which causes a homozygous C-to-T transition resulting in an R248W substitution in the p18 protease subunit. This leads to caspase-8 deficiency which may be associated with type IIB autoimmune lymphoproliferative syndrome and cancer due to defective apoptosis and the accumulation of mutations (Chun, et al., 2002).
A two-base pair deletion in exon 7 of the caspase-8 gene is associated with hepatocellular carcinoma. This somatic mutation may cause a frameshift which may result in premature termination of amino acid synthesis in the p10 subunit due to a frameshift (Soung, et al., 2005).
A single-nucleotide polymorphism in the Casp8 gene results in the D203H substitution which provides protection against breast cancer (Macpherson, et al., 2004; Frank, et al., 2005). However, it may be that another variant in strong linkage disequilibrium with D203H may reduce susceptibility to breast cancer (Cox, et al., 2007).
A six-nucleotide insertion/deletion polymorphism in the promoter region is associated with reduced susceptibility to lung cancer as well as esophageal, gastric, colorectal, and cervical cancer in an allele-dependent manner (Sun, et al., 2007).
Cite This Work
To export a reference to this article please select a referencing stye below:
Related Services
View allDMCA / Removal Request
If you are the original writer of this assignment and no longer wish to have your work published on UKEssays.com then please click the following link to email our support team:
Request essay removal


